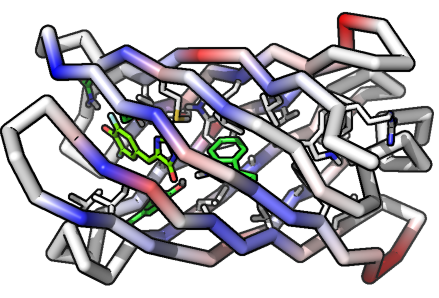
Bifurcation of High- and Low-Energy Electrons in Microbial Metabolism
Scientists at ALS-ENABLE beamline 12.3.1. used small angle x-ray scattering to understand a microbial protein involved in the bifurcation of high and low energy electrons in microbial metabolism. By analyzing the SAXS data, the SIBYLS team was able to find that NADH activates the protein’s mechanics, allowing it to act as the wheel and ropes of a pulley system enabling an unfavorable transformation by coupling to a favorable transformation or electron bifurcation. In other words, the system is able to liberate two electrons from one donor and channels them to two separate acceptors.
READ ARTICLEUnderstanding of the variability within the DHOase family of proteins
ALS-ENABLE beamline scientist, Jay Nix, contributed to this paper reporting the structural analysis of dihydroorotase (DHOase) from the hyperthermophilic and barophilic archaeon Methanococcus jannaschii. DHOase catalyzes the reversible cyclization of N-carbamoyl-l-aspartate to l-dihydroorotate in the third step of de novo pyrimidine biosynthesis. DHOases form a very diverse family of enzymes and have been classified into types and subtypes with structural similarities and differences among them. This is the first archaeal DHOase studied by x-ray diffraction. Its structure and comparison with known representatives of the other subtypes help define the structural features of the archaeal subtype. The M. jannaschii DHOase is found here to have traits from all subtypes. Contrary to expectations, it has a carboxylated lysine bridging the two Zn ions in the active site, and a long catalytic loop. It is a monomeric protein with a large β sandwich domain adjacent to the TIM barrel. Loop 5 is similar to bacterial type III and the C-terminal extension is long.
READ MORE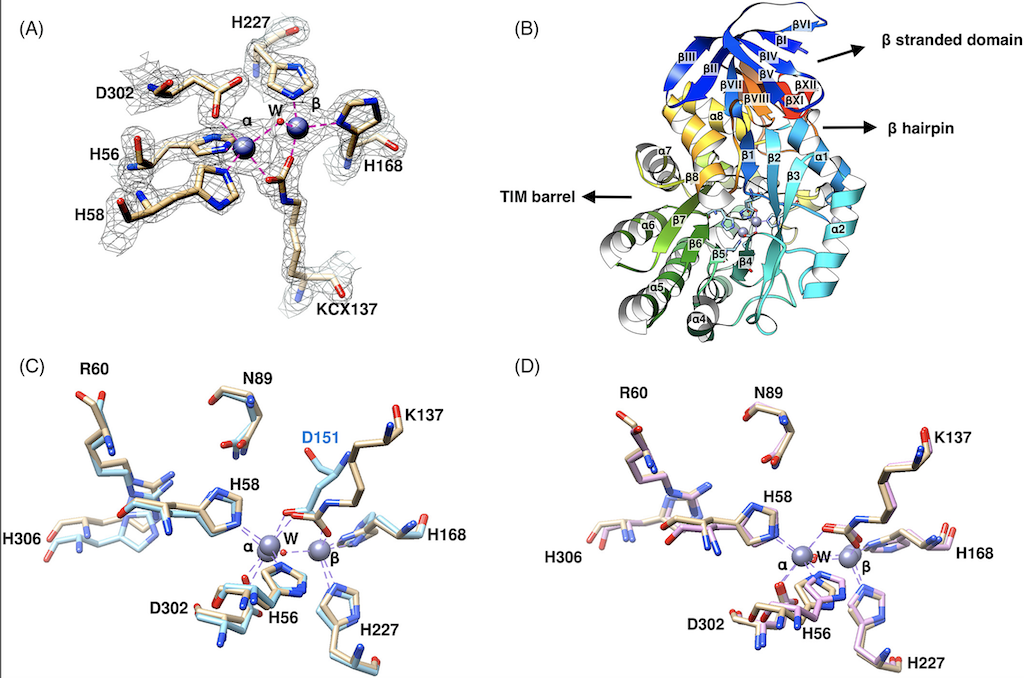
X-ray footprinting used for antibody characterization
Our review made the cover! This article describes how the method of hydroxyl radical protein footprinting, a solution-state method, can provide structural and kinetic information on antibodies or antibody–antigen interactions useful for therapeutic antibody design.
LEARN MORE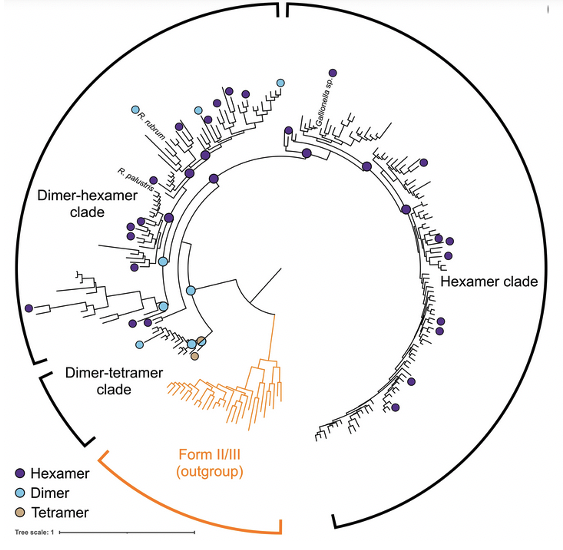
Evolution and Innovation of RuBisCO
ALS-ENABLE beamline scientists Daniel Rosenberg and Michal Hammel collaborated with the Shih Lab at UC Berkeley to study patterns of oligomerization in rubisco, nature’s most abundant carbon-fixing enzyme. By using SEC-SAXS-MALS, they were able to conduct a diversity-driven structural study across extant and ancestral form II rubisco and retrace the evolutionary trajectory taken during the evolution of the enzyme.
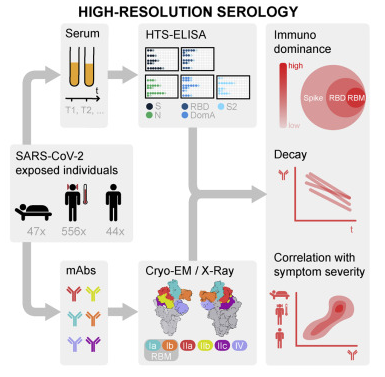
PRESS RELEASELINK TO PUBLICATIONStructural basis of SARS-CoV-2 Omicron immune evasion and receptor engagement
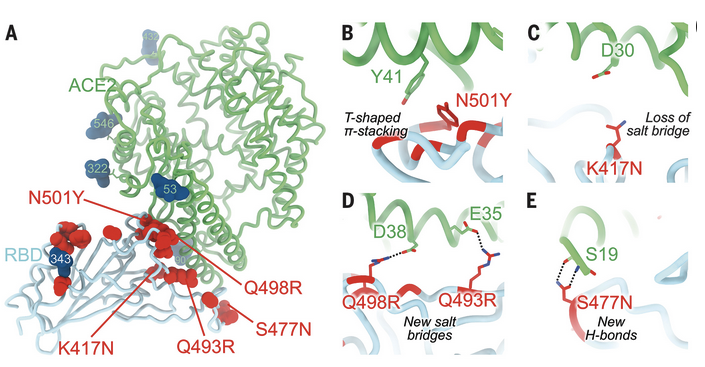
Two years into the COVID-19 pandemic, several highly infectious variants of SARS-CoV-2 have emerged. To date, the most potent of them is the Omicron variant. In this paper, ALS-ENABLE scientist Dr. Jay Nix (BL 4.2.2) along with others shed light on the Omicron antigenic shift.
READ MOREThe figure to the right shows the molecular basis of human ACE2 recognition by the SARS-CoV-2 Omicron RBD.
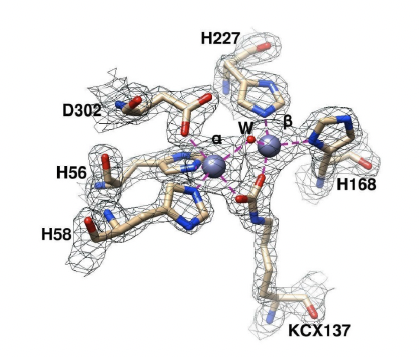
The active site of M. jannaschii DHOase superimposed on a 2mFo-DFc map contoured at 1.7σ.
Crystal Structure of an Archaeal Dihydroorotase
Dr. Jay Nix at ALS-Enable beamline 4.2.2 collected crystallographic data for this first X-ray diffraction study of the archaeal DHOase, a zinc metalloenzyme that functuions in the pyrimidine nucleotide biosynthesis. Dr. Nix was also key in determining the structure, finding both similarities and differences from other known DHOases.
READ MOREIdentifying potential new drugs for SARS-CoV-2
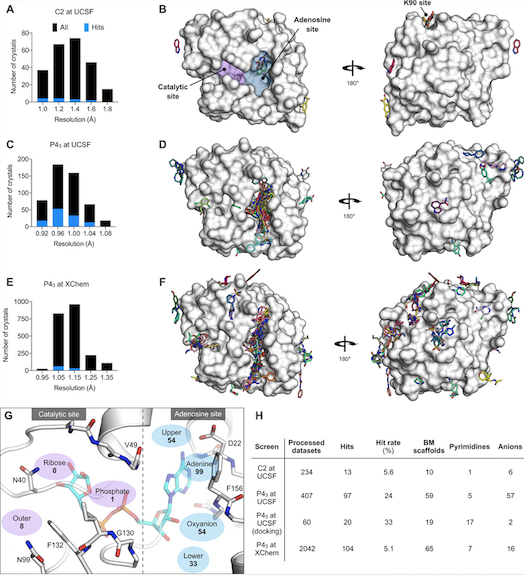
Crystallographic data collected at ALS-ENABLE beamline 8.3.1 was central in this study identifying small chemical fragments that bind to the Nsp3 macrodomain (Mac1) of the SARS-CoV-2 virus. This protein interferes with immunity, so a drug that inhibits it (sticks to it and stops it from doing that) will help the immune system win. By looking at the 3D image of all the bound fragments at once they came up with a new, larger molecule that sticks tighter than any of the fragments alone. Along with computational docking efforts, this result lays a road map for making even better inhibitors that stick even tighter, minimizing potential side effects.
LEARN MOREUsing SAXS to evaluate radical scavengers
SIBYLS (beamline 12.3.1) user Tim Stachowski and the Snell group make the cover of the Journal of Synchrotron Radiation for their paper describing their work using Small-angle X-ray Scattering to evaluate how three scavengers commonly used in crystallographic experiments perform in solution at 10°C. In this study, they found the SAXS-based method can detect damage at X-ray doses far lower than those accessible crystallographically, thereby providing a detailed picture of scavenger processes. Their approach provides a platform for more systematic and comprehensive screening of radioprotectants that can directly inform mitigation strategies for both solution and crystallographic experiments, while also clarifying fundamental radiation damage mechanisms.
READ MOREA study of antibodies that inhibit noroviruses
A study using crystallographic data collected at beamline 5.0.1 works towards the development of a vaccine to combat an acute gastroenteritis disease caused by a human norovirus. Since 2010, according the Center for Disease Control, an estimated 19 to 21 million people in the United States annually have been affected by this disease. Worldwide, there were 1.8 billion cases, 18% of which were caused by HuNoVs.
Learn More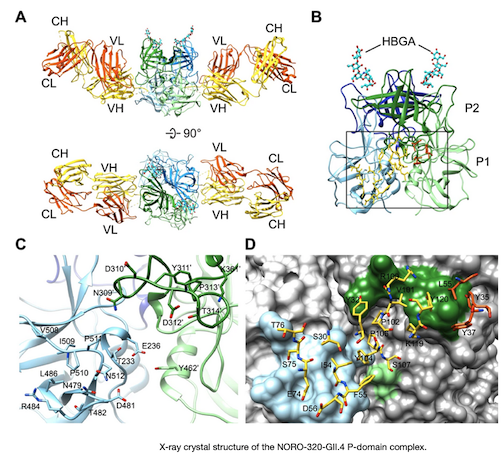
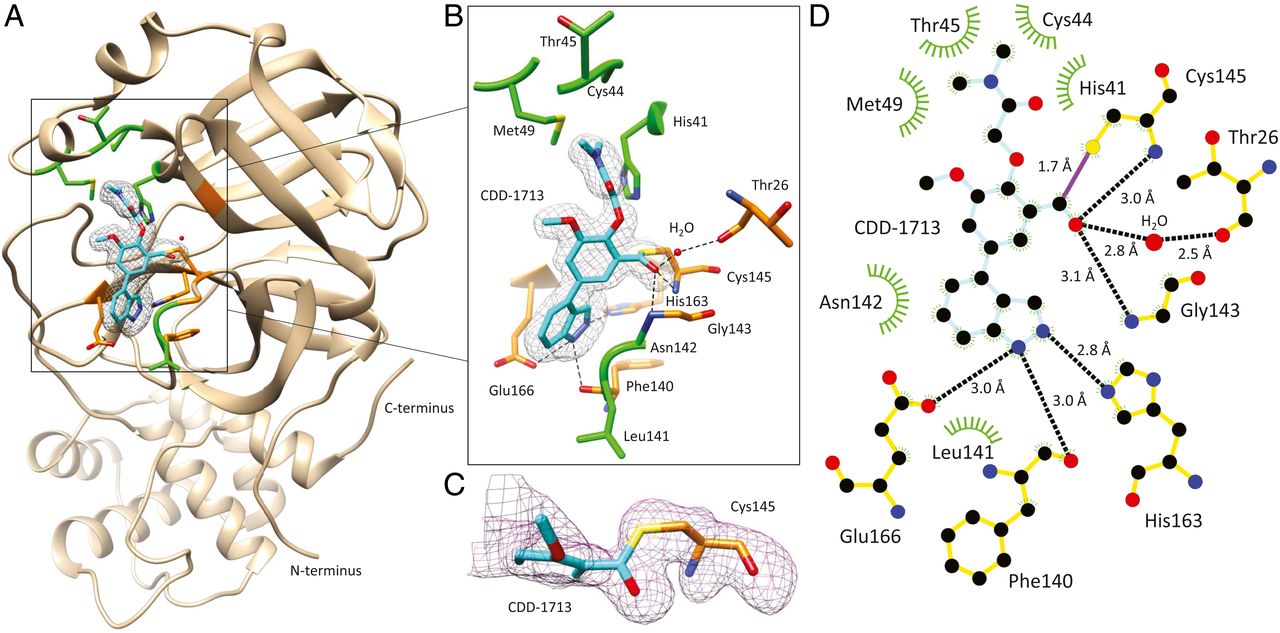
MX yields access to SARS-CoV-2 Mpro inhibitors
Collaborative Crystallography beamline 5.0.1 used an affinity-based screen of 4 billion DNA-encoded molecules en masse to identify a potent class of virus-specific inhibitors of the SARS-CoV-2 main protease (Mpro) without extensive and time-consuming medicinal chemistry to help combat the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) that has, so far, killed more than 4 million humans globally.
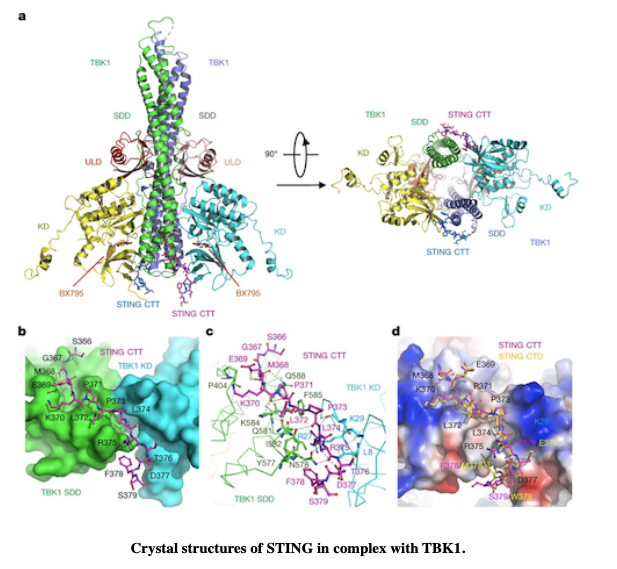
LEARN MOREA conserved PLPLRT/SD motif of STING mediates the recruitment and activation of TBK1
Crystal structures from diffraction data collected at beamlines 5.0.2 and 8.2.2 provide a structural basis for the development of STING agonists and antagonists for the treatment of cancer and autoimmune disorders.
LEARN MOREStructural biology in the spotlight
ALS Enable scientist helps to determine the atomic structure of a coronavirus protein
Using X-ray crystallography at beamline 5.0.2, Marc Allaire, helped to build an atomic model of ORF8, laying the groundwork for other groups to study the protein in more detail.
READ MORESIBYLS makes the cover of Protein Science
SIBYLS scientists Michal Hammel and John Tainer describe the mechanistic insights into NHEJ structural biochemistry determined by small angle X-ray scattering (SAXS) data collected at beamline 12.3.1.
SAXS results were combined with with X-ray crystallography (MX) and cryo-electron microscopy (cryo-EM).
The cover
The cover image, like the old car engine malfunctioning without a linchpin and flexible belt, shows the NHEJ efficiency arises from a flexible conformational control principle. The NHEJ components were matched on the SKODA 100 car engine built in 1973 in Czechoslovakia.
READ MOREprobing real-time solvent accessibility
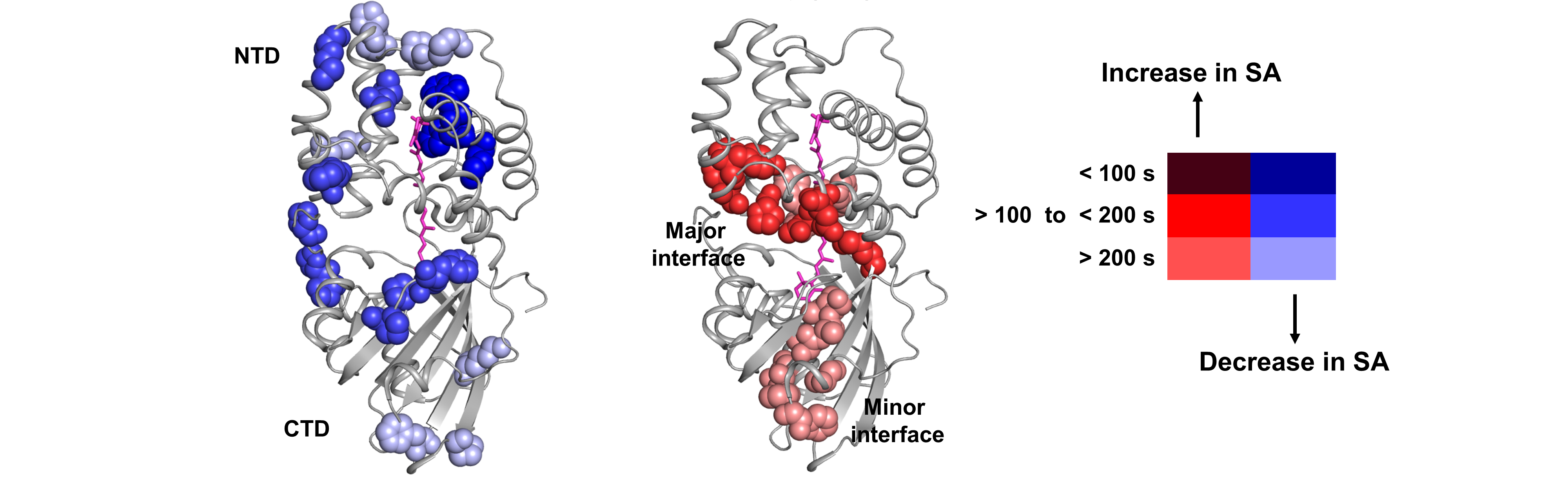
X-ray footprinting
X-ray footprinting of the Orange Carotenoid protein shows the various regions of solvent accessibility changes upon photoactivation of the protein.
Learn more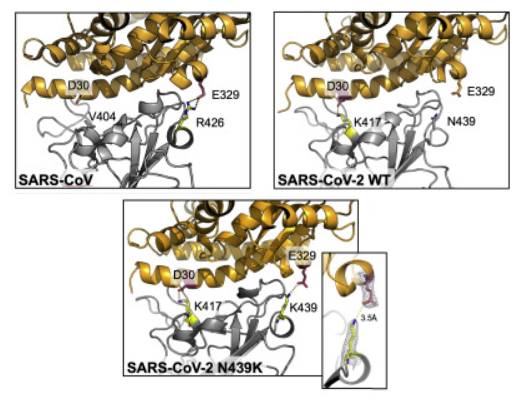
ALS-ENABLE scientist contributes to Cell publication on SARS-CoV-2 spike protein
ALS-ENABLE scientist, Jay Nix at the MBC beamline 4.2.2 participated in the structure solution for a recently published paper in Cell, Circulating SARS-CoV-2 spike N439K variants maintain fitness while evading antibody-mediated immunity
LEARN MORE
Nasal spray prototype
Helping to stop the rise of SARS-CoV-2
Scientists at ALS Enable contribute to groundbreaking research that may help to stop the rise of SARS-CoV-2 infections that have had devastating effect on our health and economy worldwide. In the article (currently in preprint) the authors describe the development of single-domain antibodies (nanobodies) that potently disrupt the interaction between the SARS-CoV-2 Spike and ACE2. This discovery may enable aerosol-mediated delivery of this potent neutralizer directly to the airway epithelia, promising to yield a widely deployable, patient-friendly prophylactic and/or early infection therapeutic agent to stem the worst pandemic in a century.
LEARN MOREGuiding the design of COVID-19 vaccines and therapeutics
ALS Enable scientists contribute to an article published in CELL (volume 183, issue 4, November 12, 2020) in which the authors describe the analysis of the specificity and kinetics of neutralizing antibodies (nAbs) elicited by SARS-CoV-2 infection. This discovery is a crucial part of understanding immune protection and identifying targets for vaccine design.
Link to PublicationFighting bacterial infections
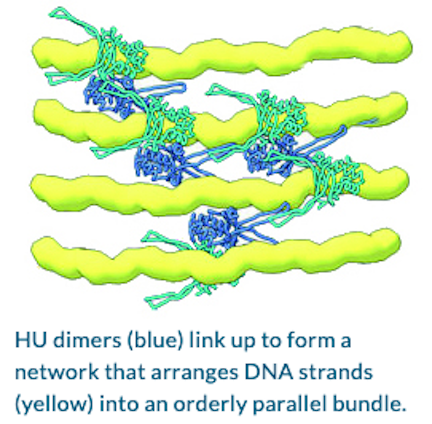
How Proteins Remodel DNA in Bacteria under Stress
ALS-ENABLE beamlines use multiscale, multimodal visualization techniques to clarify how proteins remodel bacterial DNA in response to stressful environments. At the mesoscale, small-angle x-ray scattering (SAXS) at Beamline 12.3.1 made it possible to determine the overall shapes of HU–DNA complexes in solution. At the nanoscale, protein crystallography at Beamline 8.3.1 allowed identification of the molecular-level mechanisms that mediate DNA bundling.
READ MORE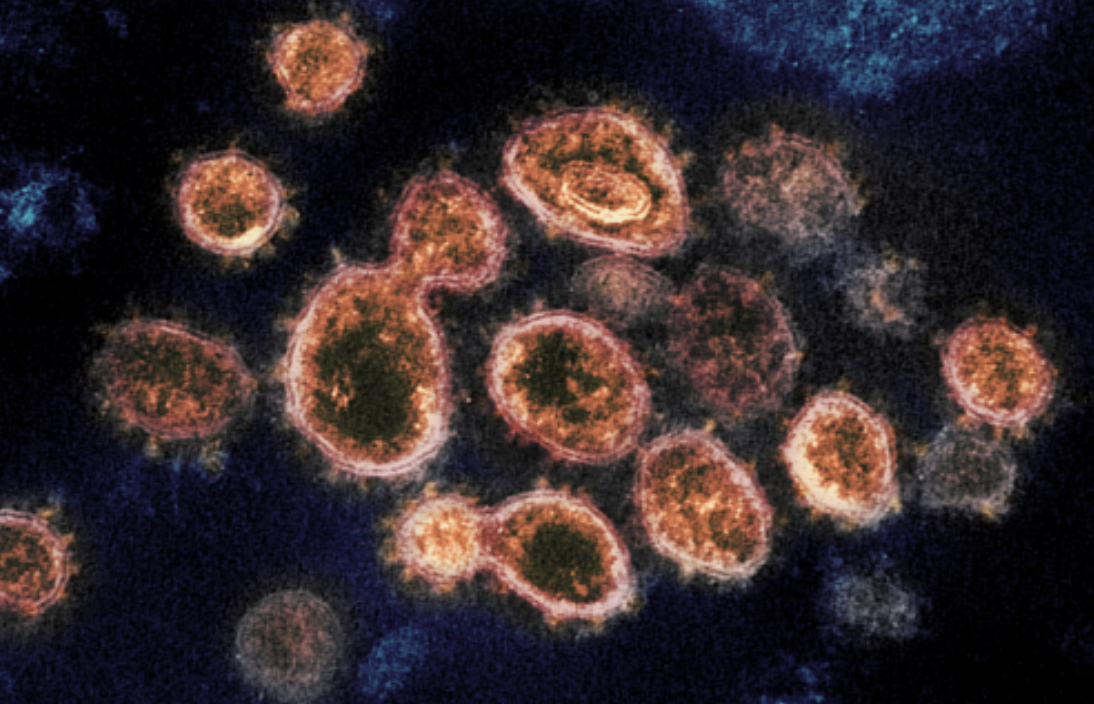
X-ray Experiments zero in on COVID-19 Antibodies
With help from ALS-ENABLE beamline 5.0.2, an international scienctific team has identified an antibody that neutralizes the COVID-19-causing coronavirus. In a paper published in Nature, the scientists note that the antibody is already moving toward clinical trials.
LEARN MOREAssembly Lines for Designer Bioactive Compounds
Researchers successfully bioengineered changes to the molecular “assembly line” for bioactive compounds, based in part on insights gained from Small-Angle X-ray Scattering (SAXS) data collected at SIBYLS beamline 12.3.1.
LEARN MORE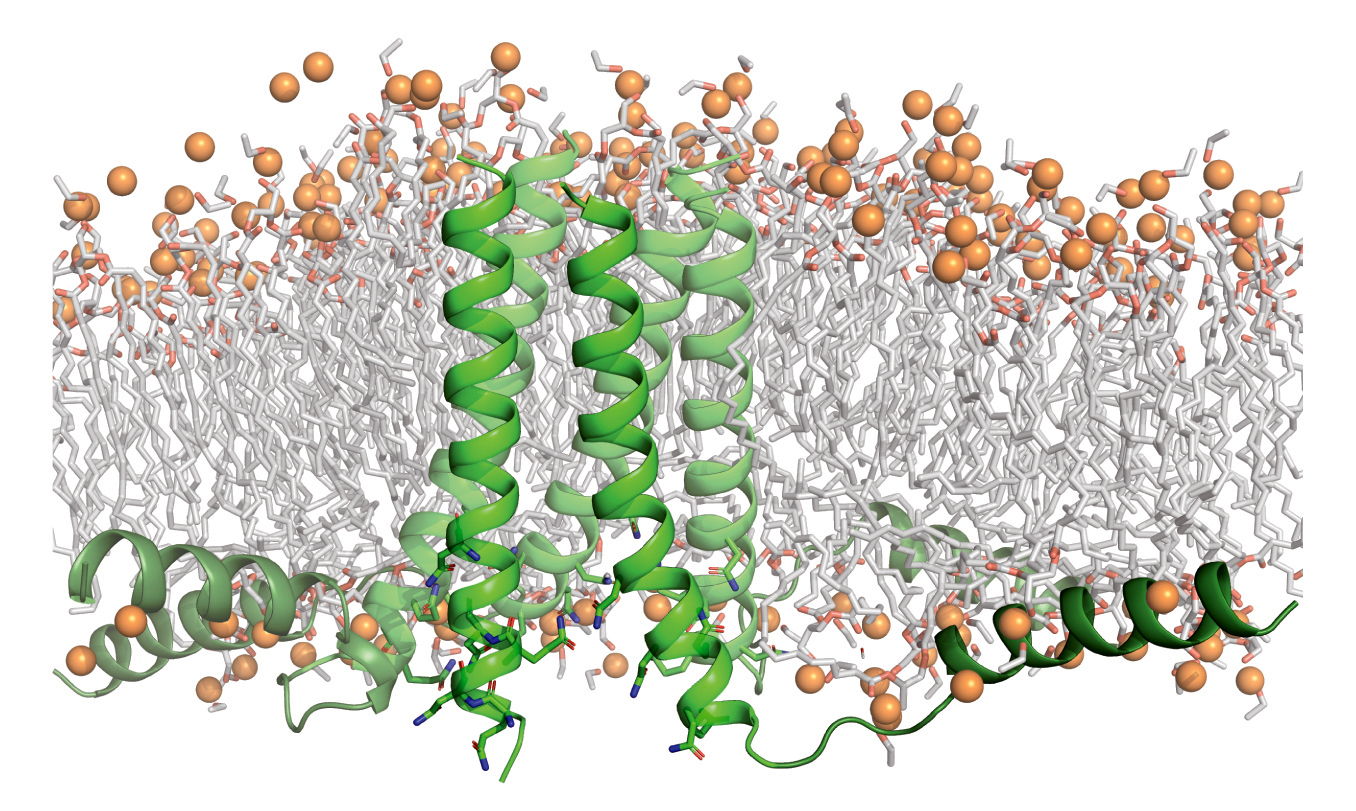
Characterizing the forces that hold membrane proteins together
Designed membrane proteins are used to better understand the competing forces that stabilize this category of protein.
LEARN MOREDesigner proteins that fit together like DNA
Proteins designed to mimic the structure of DNA can be used as building blocks for self-assembling protein machines.
Read MoreToward defeating influenza
Structures of several influenza antiviral drug molecules bound to their targets in both open and closed conformations provide an atomic-level blueprint from which to design more effective anti-influenza drugs that can overcome growing drug resistance.
Learn moreDesigning a protein to capture small molecules
For the first time, scientists have created, entirely from scratch, a protein capable of binding to a small target molecule.
LEARN MORE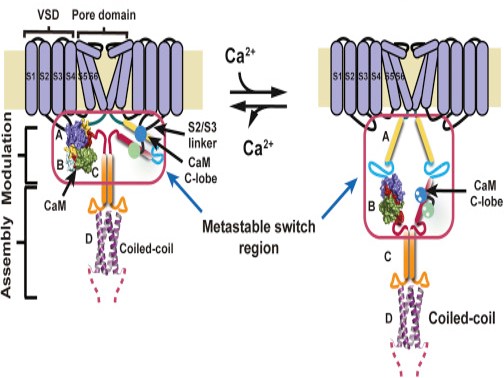
Decoding a Calcium dependent switch
Certain voltage-gated proteins in cell membranes are responsible for controlling the flow of ions across the membrane and thus are important in electrical activity in the heart and brain. This study using both diffraction and scattering beamlines of ALS-ENABLE revealed the role of calcium in the regulation of a class of these important proteins.
Read MoreHow antidepressants block serotonin transport
Serotonin is a diminutive and deceptively simple-looking neurotransmitter molecule, yet a very complex “machinery” is required for neurotransmitter recognition, transmission, and recycling. The malfunctioning of this protein machinery can cause conditions such as depression, obsessive-compulsive disorder, aggression, anxiety, and Parkinson’s disease. In addition, drugs of addiction, such as methamphetamines, act on this system, causing serious damage and profoundly affecting the well-being of individuals. At the ALS, researchers were able to obtain x-ray crystallographic structures of the difficult-to-crystallize human serotonin transporter bound to two commonly prescribed antidepressant drug molecules. The resulting details about the transporter structure and mechanism will help in the design of new, more effective therapeutics for treating depression and anxiety.
Read MoreBehind the new gene-editing tool
The CRISPR-Cas system has recently transformed the way that scientists are able to perform gene editing, making it much easier and more accurate than ever before. In this study, the structures of the system with DNA substrates revealed just how specific DNA code is located with such specificity.
Read More